DNA Under The Microscope
Electron & Atomic Force Microscopy
Overview
DNA (Deoxyribonucleic acid) is the molecule that contains within it all the instructions and information about an organism. This is to say that DNA contains information regarding how the organism will develop, how it lives and reproduces etc. Therefore, the DNA may be described as the blueprint of a living organism.
Given that DNA molecules are found inside the cells, they are too small to be seen with the naked eye. For this reason, a microscope is needed. While it is possible to see the nucleus (containing DNA) using a light microscope, DNA strands/threads can only be viewed using microscopes that allow for higher resolution.
Microscopy
To view the DNA as well as a variety of other protein molecules, an electron microscope is used. Whereas the typical light microscope is only limited to a resolution of about 0.25um, the electron microscope is capable of resolutions of about 0.2 nanometers, which makes it possible to view smaller molecules. This is achieved because electron microscopes use electron beams rather than the visible light used for light microscopes.
DNA Electron Microscopy
Requirements
- Electron microscope
- Heavy metal salts (Lead perchlorate, uranyl acetate, lanthanum nitrate)
- DNA sample (nucleic acids)
- Formaldehyde
Procedure
For this procedure, the steps involved:
- Spraying DNA onto a grid with a glass nebulizer (using freshly cleaved mica is advised)
- Add latex spheres - Latex spheres or ferritin used here serve as reference particles
- Spread the DNA solution/preparation and protein (1:10 to 1:100) on water surface in a Langmuir trough - This is known as the Kleinschmidt's technique. It is used for spreading nucleic acid in protein to form a film of protein that retains DNA on the surface
Once the preparation is ready, it is ready for staining
Staining Procedure
- The nucleic acid is applied to the grids and immersed in the staining solution for between 10 minutes and 4 hours.
- Rinse the grids using redistilled water at a pH of 6.0 three times and view under the microscope
- In the event that the stained grid will not be viewed immediately, then they can be stored in an evacuated desiccators over phosphate P205
Observation
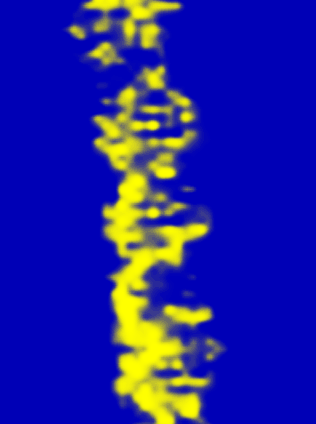
Using the heavy salts allow for higher contrast that makes it possible to view single molecules of the DNA
Scanning Transmission Electron Microscope (STEM)
STEM microscopy has been shown to operate in a dark-field mode thus providing high contrast of biological molecules. Because of its dark-field images, this technique has also been shown to have a great advantage in that it allow for direct visualization of unstained strands of DNA.
Through the high contrast provided, the technique also makes it possible for researchers to be able to identify any problems with the sample. The procedure for this technique is a lot similar to typical electron microscopy for DNA. However, through STEM, researchers obtain both mass and structural information of single-stranded DNA.
Cryo-Electron Microscopy
Cryo-electron microscopy is one of the techniques that have been shown to be particularly successful in revealing the structure of DNA.
Unlike the transmission electron microscope, Cryo-EM uses frozen samples and electron beams that are gentler to view the sample. This allows researchers to view biological molecules without causing any damage to them in the process.
For this technique, a small amount of the sample in solution is first applied to an EM grid similar to the process used to view DNA strand under electron microscope (EM). The grid with the thin layer is then immersed in liquid ethane (at -180 degrees c) to trap the molecules in water crystal/ice. This ensures that the sample remains is not destroyed when being viewed under the microscope.
Here, it is worth noting that samples prepared for this technique (Cryo-EM samples) tend to be highly sensitive to electron damage. For this reason, low electron doses of about 10–20 e−/Å2 are used to ensure that the sample is not damaged.
While the sample is exposed to low doses of the electrons, the layer of ice around the sample also helps protect the sample during the process.
Cryo-Electron Tomography (CET)
Through recent advancements in this technique, researchers were able to develop an improved technique of Cryo-EM known as Cryo-electron tomography (CET). Using this technique, it has become possible for researchers to develop 3D structures of various proteins and DNA strands.
Essentially, the process involves the capture of many images of the sample from various angles and using the images to build a 3D structure. Using this technique, researchers have managed to develop and present a variety of 3D images of DNA strands showing the structure of DNA from different angles.
Atomic Force Microscopy
Apart from the electron microscope techniques used to study DNA, methods such as Atomic Force Microscopy (AFM) are also being used for the same purpose. Using this technique, it has become possible for researchers to measure the length of these strands.
Requirements (for AFM)
For this technique, some of the materials required include:
- DNA samples (such as X174 virion DNA)
- AFM microscope
- Freshly-cleaved ruby mica circle
- Magnesium acetate
- Water
- AC glow-discharge
- Formaldehyde
Procedure
- For single stranded DNA, the procedure involves the following steps:
- Soak freshly-cleaved ruby mica circles in 33 mM of magnesium acetate for between 4 and 24 hours
- Sonicate in Millipore water for about 5 minutes to remove any excess magnesium acetate
- Use compressed air for drying
- Expose to AC glow-discharge for about 20 seconds in 100 militorr air
- Almost immediately invert the circles in about 7 micro liter drops of the DNA strands (singles stranded) in 0.5 percent formaldehyde and 15 mM ammonium acetate - This is deposited to a parafilm
- Allow to stand for about 4 minutes and rinse the mica using about 3 drops of water
- Dry using compressed air
- If it is not to be used immediately, store the preparation in a desiccator on drierite
For Double stranded DNA, the same procedure is used but formaldehyde and ammonium acetate is not used.
Imaging
- During imaging (AFM-imaging) the process was carried out under 100 percent propanol using either Nanoscope II or III AFM. Here, an O-ring was used to apply the propanol by laying it on the sample in the device. The process involved:
- Placing a drop of alcohol on to the cantilever in the fluid cell
- Positioning the cantilever over the O-ring and firmly clamped
- Ensuring that the amount of fluid was sufficient (this was determined by ensuring that no bubbles remained on the light path)
During imaging, the minimum force was used. This ensured that the cantilever did not lift the sample.
To measure the length of DNA strands using this technique, the images are first enlarged and a fine chain laid along the DNA contours. However, a better way of making the measurements has been shown to involve direct measuring of the top-view of the DNA images in the nanoscope. This method simply involves summing up given points in the image.
Scanning Tunneling Microscopy (STM)
The STM can also be used to view DNA molecules. It is capable of imaging objects at atomic levels, which makes it a good tool for viewing DNA molecules.
For this technique, several methods may be used to prepare the sample for imaging, these include:
Method #1
- Place a droplet of the DNA solution on the substrate
- Allow it to dry in air (air dry)
- Contain the solution in 10mM potassium chloride to stabilize the DNA
- Sonicate the solution using a
- Biosonik IV probe sonicator for about30 minutes at 160 W
- Imaging - Imaging using STM can be directly carried out on the layer
Method #2 - This method is similar to the first method, but involves vacuum drying the solution and containing it in 10mM ammonium acetate. Sonication was also skipped.
Method #3
- Place 10 micro liters of distilled water on a piece of parafilm - Water causes the phage to burst releasing DNA
- Add 10 micro liters droplet of lysed DNA particles (lysed T7 bacteriophage)
- Briefly place a substrate on the droplet to pick up the lysed particles
Following imaging, it is possible to identify the DNA through their heights and width. Here, DNA strands can be clearly seen arranged parallel to one another.
One of the most recent methods slightly deviates from the others and involves dissolving DNA in an aqueous solution and placing it in a layer of highly oriented graphite before allowing it to air dry. Here, tungsten-wire tips are used while scanning is carried out under atmospheric conditions. Scanning produces high resolution images of the double helix DNA.
Wolfgang Schonert, original author at GSI.de
http://web-docs.gsi.de/~bio/RESEARCH/rastermicro.html
DNA Structure
As mentioned, DNA (deoxyribonucleic acid) carries within it genetic, hereditary material and resides in the cell nucleus of every organism. Essentially, the strands of DNA are composed of repeated patterns of six molecules which include; deoxyribose (a five carbon sugar) a phosphate group as well as four nitrogenous bases (cytosine (C), thymine (T), adenine (A) and guanine (G)).
The linear order of the bases represents one of the most important features of DNA given that their paring allows for important encoding of information required for development and life of the organism. The structure of DNA (double-helical) is of great significance given that it serves to protect the base atoms.
* Nucleotides are the basic units of DNA. Each nucleotide is composed of a single molecule of sugar, a single phosphate molecule and one nitrogenous base.
See Also:
What is Near Field Scanning Optical Microscopy?
Return from DNA under the Microscope to MicroscopeMaster Home
References
Walther Stoeckenius. Electron Microscopy of DNA Molecules "Stained" With Heavy Metal Salts .
Hele G.Hansma Rober L.Sinsheimer Min-Qia Li and Pau K.Hansma (1992) Atomic force microscopy of single and double-stranded DNA.
http://iopscience.iop.org/article/10.7567/JJAP.56.08LB02
http://cs.boisestate.edu/~amit/teaching/342/lab/structure.html
Find out how to advertise on MicroscopeMaster!
![DNA structure by Zephyris (Own work) [CC BY-SA 3.0 (https://creativecommons.org/licenses/by-sa/3.0) or GFDL (http://www.gnu.org/copyleft/fdl.html)], via Wikimedia Commons DNA structure by Zephyris (Own work) [CC BY-SA 3.0 (https://creativecommons.org/licenses/by-sa/3.0) or GFDL (http://www.gnu.org/copyleft/fdl.html)], via Wikimedia Commons](https://www.microscopemaster.com/images/492px-DNAStructureKeyLabelled.pnNoBB.png)