Gram Positive and Gram Negative Bacteria
Structures/Characterisitics
Overview
To identify Gram positive and Gram negative bacteria a number of approaches are used to classify/identify organisms. In clinical microbiology, phenotypic typing schemes (particularly Gram staining) are some of the most common and effective techniques for bacteria identification.
Discovered in 1884, Gram stain technique has proven to be one of the most useful phenotypic classification systems through which bacteria can be identified based on their staining properties and general morphology. Using this technique, bacteria are classified as either Gram positive or Gram negative.
The difference between the two groups is as a result of a number of chemical/surface structures that include:
- Teichoic Acids
- Lipopolysaccharides
- Peptidoglycans
- Lipoteichoic Acids
* While Gram positive and Gram negative bacteria are of clinical significance, not all bacteria in these groups are harmful/cause disease.
Bacteria
Bacteria are single-celled prokaryotes ubiquitous in nature. As such, they can be found in different types of environments (ranging from aquatic to the hot rocks of the Mojave Desert).
In these environments, bacteria may exist as free-living organisms (making their own food or feeding on dead organic matter etc), in symbiosis with other organisms (e.g. Zoamastogopera found in the digestive system of termites) or as parasitic organisms that can cause harm and disease in their hosts.
* Bacteria are thought to have been the first living organisms on earth.
* While bacteria can cause disease, they are also useful in medicine, food and other industries.
* Bacteria are neither plants nor animals.
Some of the main functions of bacteria (in general) include:
· Can reproduce asexually (binary fission) and asexually (transduction and conjugation etc) - They can also form spores
· Being prokaryotes, they lack membrane-bound organelles
· Vary in shape and can, therefore, be classified based on their distinct shapes (Bacilli, Cocci, Spirilli)
· Their wall consists of peptidoglycan
· Some bacteria move using a tail-like structure known as flagella
Gram Positive Bacteria
Gram positive bacteria are a group of organisms that fall under the phylum Firmicutes. However, a few species have a Gram negative cell wall structure.
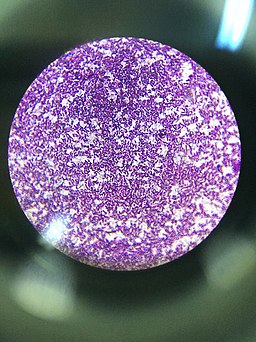
As compared to Gram negative bacteria, this group of bacteria is characterized by their ability to retain the primary stain (Crystal violet) during Gram staining (giving a positive result).
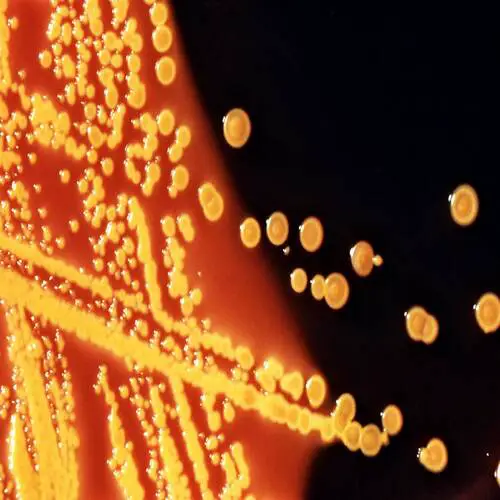
Some examples of Gram positive bacteria include:
Staphylococcus (e.g. Staphylococcus aureus) and Streptococcus bacteria (e.g. Streptococcus pyogenes) - are cocci bacteria (spherical) responsible for a number of diseases in human beings
Whereas Staphylococcus aureus can cause such pyogenic infections as septic arthritis, Streptococcus pyogenes are responsible for such infections as cellulitis, tonsillitis, and rheumatic fever.
Compared to some of the bacilli bacteria, bacteria are non-motile (lack motility organs such as flagella) and do not form spores.
Gram positive bacilli - include such bacteria as Clostridia, Bacillus, and Listeria among others. While some of the species are characterized by their ability to produce spores (e.g. Clostridia), others like Listeria do not. In human beings, these organisms are responsible for a number of diseases
For instance, Bacillus anthracis has been shown to cause anthrax in human beings (rare in human beings but common in animals).
* Some of Gram positive bacilli use flagella for movement (e.g. Listeria monocytogenes).
Characteristics of Gram Positive Bacteria
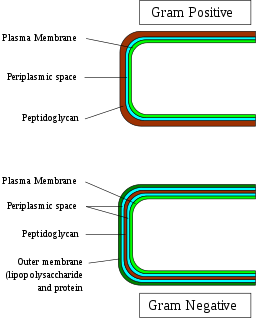
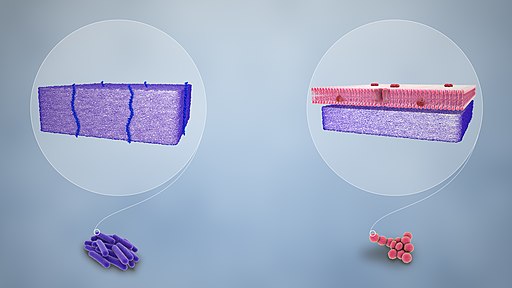
Cell Wall
The cell-wall of Gram positive bacteria consists of a number of important components that include:
Peptidoglycan
Located outside the cytoplasmic membrane of most bacteria, the peptidoglycan is one of the main components of the bacterial cell wall. For the most part, it serves to maintain cell integrity of the organism by withstanding turgor pressure.
Moreover, it contributes to the general shape of a cell in addition to providing support for various components of the cell. In Gram positive bacteria, the peptidoglycan makes up between 50 and 90 of the cell wall (between 20 and 80 nm in thickness). By enclosing the plasma membrane, it provides a protective layer that protects the cell from lysis.
Main components of peptidoglycan include:
· Glycan chains consisting of N-acetylglucosamine and N-acetylmuramic acid linked by β-1,4 bonds.
· Peptide chains - in peptidoglycan are linked to N-acetylmuramic acid by covalent bonds
* Apart from the peptidoglycan layers, the cell wall of Gram positive bacteria has also been shown to contain polysaccharide molecules. Some of the best examples of these accessory polymers include C polysaccharides found in the cell wall of streptococci bacteria and peptidoglycolipids and glycolipids found in the cell wall of Mycobacteria.
* Depending on the species, peptidic chains may vary in compositions. For instance, whereas the peptide chains are directly cross-linked through short amino-acid chains, others tend to be indirectly cross-linked.
* The peptidoglycan is in a dynamic state over the life span of the bacteria.
Teichoic Acid
Apart from the peptidoglycan layers, the cell wall also consists of molecules known as teichoic acids (polyol phosphate polymers). Here, these molecules form layers that run perpendicularly to the peptidoglycan sheets. While they are only found in Gram positive bacteria, teichoic acids are only found in some cells of this group.
Generally, they are composed of ribitol/glycerol that are joined to each other by phosphodiester bonds. Some of the other components of these molecules include glucose, N-acetylglucosamine, N-acetylgalactosamine, and D-alanine. However, the amount/level of these substituent groups varies from one group to another.
Apart from being covalently linked to the peptidoglycan layers, the teichoic acids may also be anchored to the plasma membrane located below the peptidoglycan layer.
While the main function of these molecules are not well understood, studies have shown the phosphodiester bonds linking the monomers of teichoic acid to result in the overall negative charge of the cell wall of Gram positive bacteria.
* While the function of teichoic acids is not fully understood, researchers have suggested that they are involved in the regulation and arrangement of muramic acid sub-units. It has also been associated with antigenic properties that activate given immune cells.
Lipids
In the cell wall of Gram positive bacteria, lipid content makes up less than 5 percent of the cell wall. Here, these molecules play the role of anchoring the membrane.
Gram Negative Bacteria
Like Gram positive bacteria, Gram negative bacteria are also well distributed in different environments across the globe. Whereas E. coli among other Gram negative bacteria can be found in the gastrointestinal tract, a number of species can be found in marine environments.
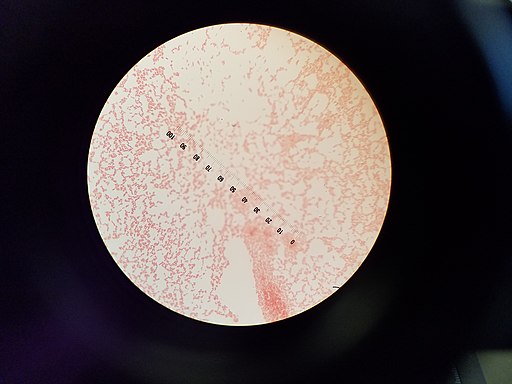
Compared to Gram negative bacteria, these bacteria are do not retain the primary stain and thus appear red when viewed under the microscope.
Some examples of Gram negative bacteria include:
· Gram negative cocci - Some of the most common Gram negative cocci bacteria include members of the genus Neisseria (e.g. N. meningitidis and N. gonorrhea) and Moraxella (e.g. M. catarrhalis). In human beings, these bacteria are responsible for such diseases as meningitis (caused by N. meningitidis) and bronchopneumonia (caused by M. catarrhalis).
· Gram negative coccibacilli - As compared to Gram negative cocci bacteria, Gram negative coccibacilli are short rods (intermediates spherical cocci and bacilli rods). They include a number of species responsible for human diseases including; Haemophilus influenzae that cause ad pneumonia, Haemophilus ducreyi that causes genital ulceration and Acinetobacter species capable of causing such wound diseases among other diseases.
· Gram negative bacilli - There are different groups of Gram negative bacilli bacteria including those described as Anaerobic or Aerobic. Apart from E. coli, some of the other bacteria classified as Gram negative bacilli include members of the genera Fusobacterium and Bacteroides.
In human beings, they are responsible for a number of human diseases including oral pleuropulmonary and infections in the female genital tract among others.
* According to studies, Gram negative bacteria, led by E. coli, make up over 70 percent of urinary tract infections. See Urine Analysis
Characteristics of Gram Negative Bacteria
As with Gram positive bacteria, Gram negative bacteria also contain the peptidoglycan polymer in their cell wall. While this polymer is thin (2 to 4 nanometers in thickness with just about 3 layers of peptidoglycan) in Gram negative bacteria, it's also composed of long glycan strands that are cross-linked by peptide molecules.
This composition serves a number of functions including protecting the bacterial cell from lysis.
Development of peptidoglycan in Gram negative bacteria
In Gram negative bacteria, the development of peptidoglycan has been shown to start with the polymerization of glycans. Here, the carbohydrate-based polymer is polymerized to form strands consisting of about 100 subunits of disaccharide.
In E. coli, the length distribution of the strands is broad (consisting of between 20 and 30 units of disaccharides). However, this is largely dependent on a number of factors including environmental conditions, bacterial strains, and the growth phase among a few others.
In some of these bacteria, this length may increase to over 80 disaccharide units. The disaccharides themselves are covalently linked to peptides (wound around the glycan strands) which may also be cross-linked to those emerging from other glycan strands.
* In E. coli, over 80 percent of the peptidoglycan exists as a monolayer.
* Peptidoglycan makes up about 10 percent of the cell wall in Gram negative bacteria.
Compared to Gram positive bacteria that lack the external membrane, the peptidoglycan layer in Gram negative bacteria is surrounded by an outer membrane. For this reason, the peptidoglycan layer is not the outermost part of the cell wall in these organisms.
While it's similar to the plasma membrane in several respects, the two are different in a number of ways. For instance, compared to the plasma membrane of the cell (inner membrane), Lipid A of the Lipopolysaccharides, along with phospholipids produce the outer leaflet of the bi-layer structure of the outer membrane.
* Apart from the lipid, the outer membrane also contains a membrane protein known as porin. Arrangement of this protein regulates the passage of molecules across the membrane.
Lipopolysaccharides
One of the most defining characteristic of Gram negative bacteria is the presence of different forms of a complex macromolecule known as lipopolysaccharide (LPS) - Also known as endotoxins in Gram negative bacteria. They are particularly important for bacteria in that they contribute to the pathogenesis of the organisms.
Lipopolysaccharide molecules consist of three main regions that include:
- Lipid A structure
- A core composed of such components as ethanolamine, galactose, and heptose among others
- Polysaccharide chains
Some of the other components of Gram negative bacteria include:
- Cytoplasmic membrane
- Mesosomes
- Cytoplasm
- Nucleoid
- Endospores
Functions of Cell Components in Gram negative bacteria
As previously mentioned, the peptidoglycan present in the cell wall of Gram negative plays an important role in preventing cell lysis (osmotic lysis). However, various components of the cell wall also have a number of other important functions for the organism.
For instance, porins present in the cell wall are assembled in a manner that creates regulative channels on the cell wall. Here, these proteins control/regulate the movement of hydrophilic molecules across the outer membrane of the bacteria, only allowing hydrophilic molecules of molecular weights below 600.
Apart from regulation of material movement across the outer membrane, the outer membrane is also involved in a number of functions including molecule signaling as well as enzyme activities among others.
In addition to these components, one of the other important components of the outer membrane in Gram negative bacteria is a slime layer that covers the membrane. By covering the surface of this membrane, this loose slime hides various components (e.g. antigens) of the membrane (which can activate cells of the immune cell).
In the process, as such, it protects the organisms in the body of the host while promoting pathogenesis. Apart from protecting the bacteria from immune cells, this substance also prevents the penetration of various drugs into the cell thus contributing to drug resistance among some species of Gram negative bacteria.
Apart from loose slime, some of the cells (both Gram negative and Gram positive bacteria) have been shown to form capsules (consisting of high molecular-weight viscous polysaccharides) on their surface. While the capsule is formed on the surface of the outer membrane in Gram negative bacteria, it's formed on the thick peptidoglycan layer in Gram positive bacteria.
In these organisms, the capsule, which can be up to 10um in thickness, encapsulated the cells promoting virulence. Like the loose slime, the capsule, which appears as a thick gel, not only hides some of the components located on the cell surface from immune cells, but also increases resistance to a number of drugs.
Gram Staining
Developed by Hans Christian Gram, Gram stain technique is one of the most commonly used methods for differentiating between Gram-positive and Gram-negative bacteria.
Essentially, this involves the use of a primary stain (e.g. crystal violet) as well as iodine solution to produce an iodine-dye complex in the cytoplasm of the bacteria that makes it possible to differentiate between the two types of cells following decolorization.
In a clinical setting, this is particularly important in that it makes it possible to not only differentiate between bacterial species, but also for diagnosing an infection.
Procedure
Sample preparation:
· Using a pair of gloves and a sterile loop/splint, create a thin film (smear) of the sample (e.g. sputum, cerebrospinal fluid, etc) on a clean glass slide
· Air-dry the sample and then heat fix by passing it over a flame several times (avoid overheating)
Staining procedure:
· Place the slide on a staining rack and flood with crystal violet (primary stain) for about 60 seconds
· Flood the slide with Iodine (Gram's iodine) for about 2 minutes
· Using 95 percent ethanol, gently decolorize the smear - Stop once the thinnest part of the smear appears colorless
· Gently wash the slide with water
· Flood the slide with the secondary stain (Safranin) for about a minute
· Allow the slide to air dry then view under the microscope under oil immersion
Each step in sample preparation and staining should be carefully followed in order to obtain good results. For instance, heat fixing not only denatures the cells, but also adheres them to the slide.
If not properly fixed, then the cells can be easily washed away during the staining process. Also, overheating causes damage to the structure of the cell which would affect the end results. As well, excessive decolorization can end up removing the primary stain.
Cell Wall Structure and Gram Stain
When viewed under the microscope, Gram negative and Gram positive bacteria will produce different results. Whereas Gram positive bacteria will appear purple/violet (having retained the primary stain: dye-iodine complex), Gram negative bacteria will appear red/pale because they do not retain the primary stain. These differences between the two groups of bacteria are due to the differences in their cell wall structure.
During the staining process, the primary stain (crystal violet) and Gram's Iodine combine to form an insoluble complex (crystal violet-iodine complex) inside the cell. When the dehydrating agent (95 percent ethanol) is added, it dehydrates the cell wall of Gram positive bacteria causing it to shrink (also causing the pores to close).
With the pores closed, the insoluble complex is trapped in the cell. In Gram negative bacteria, ethanol easily penetrates this surface and dissolves the lipid components of the outer membrane. As a result, the crystal violet-iodine complex is easily lost (removed) from the cell.
During the second round of staining (using the secondary stain/counterstain: safranin) Gram negative bacteria stain red because they take in the red color of the counterstain. However, Gram positive bacteria do not take up the counterstain given that the stain does not disrupt the primary stain in the cells.
Return to Bacteria- size, shape and arrangement
Return to Prokaryotes main page
Return from Gram Positive and Gram Negative Bacteria to MicroscopeMaster home
References
Frank Lowy. Bacterial Classification, Structure and Function.
Kerwyn Casey Huanga, Ranjan Mukhopadhyayb, Bingni Wena, Zemer Gitaia, and Ned S.
Wingreen. (2008). Cell shape and cell-wall organizationin Gram-negative bacteria.
Marie-Pierre Chapot-Chartier and Saulius Kulakauskas. (2014). Cell wall structure and function in lactic acid bacteria.
Omeed Sizar; Chandrashekhar G. Unakal. (2019). Gram Positive Bacteria.
Yunusa Thairu, Idris Abdullahi Nasir, Yahaya Usman. (2014). Laboratory perspective of gram staining and its significance in investigations of infectious diseases.
Links
https://accessmedicine.mhmedical.com/content.aspx?bookid=1792§ionid=120716441
https://serc.carleton.edu/microbelife/research_methods/microscopy/gramstain.html
https://www.bio.vu.nl/~microb/Protocols/General_protocols/Gram-Kleuring.pdf
Find out how to advertise on MicroscopeMaster!