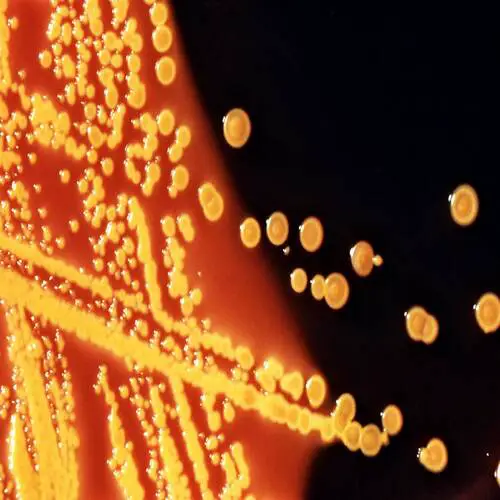
Salmonella
Classification, Causes, Microscopy, Treatment and Prevention
Overview
Salmonella includes a group of gram-negative bacillus bacteria that causes food poisoning and the consequent infection of the intestinal tract. While some of the infections can be easily treated, some of the strains have been shown to resist antibiotic treatment and are therefore deadly. For this reason, infections should not be underestimated.
There are two major species that include:
- S. bongori
- S. enterica
Classification
The genus Salmonella is closely related to Escherichia coli bacteria and is suggested to have diverged from the bacteria (E. coli) about 150 million years ago. As such, it has adapted and can be found in several niches in the environment.
Several methods of classification of Salmonella have been suggested so far. No single method/approach has been unanimously agreed on.
The following is one of the most recent classifications as used by the Center for Disease Control (CDC) as per recommendations by the World Health Organization (WHO):
- Domain: Bacteria - As bacteria, Salmonella are prokaryotes with a simple cell structure that lacks membrane bound organelles.
- Order: enterobacteriales - Gram-negative rods (Bacillus) that typically move by using flagella and do not form endospores/microcysts
- Family: Enterobacteriaceae - This is the only family in Order enterobacteriales and is composed of rod-shaped gram-negative bacteria.
- Genus: Salmonella
- Species: S. bongori and S. enterica
- Subspecies: S. bongori has a single subspecies referred to as Subspecies V.
The following are subspecies of Salmonella enterica:
- enterica I
- salamae II
- arizonae IIIa
- diarizonae IIIb
- houtenae IV
- indica VI
* In addition to the subspecies, there are also various serotyes of Salmonella that have been suggested to range from 2,200 to about 4,400 serotypes/serovar.
* Serotype grouping is based on cell surface antigens.
Serotypes (Kauffman Classification)
With regards to Salmonella serotypes, the bacteria has been shown to possess three types of antigen. These include antigen H (flagella antigen), antigen O (somatic antigen) and Vi (capsular). These antigens play an important role when it comes to grouping or serotyping the organisms.
- Antigens - This antigen is composed of lipopolysaccharide. Also refered to as somatic antigen, O antigen occurs on the outer membrane and is typically determined by the sugar sequence.
- H antigen - This includes the proteins that are found on the flagella of the bacteria. The H antigen occurs as either phase 1 or phase 2 (or both in some cases). While they may occur in either of these forms, the organisms have also been shown to change from one phase to the other. Currently, studies have identified well over 1800 serovars in this classification.
- Vi antigen - Vi is found in a few serovars and is a superficial antigen that overlies the O antigen. As such, it is an additional antigen found in such organisms as Salmonella typhi and Salmonella paratyphi C where it plays an important role in confirming serotype determination.
* Serotyping is conducted using a pure culture of the organisms isolated on non-selective agar. Some of the media that may be used include: Triple Sugar Iron (TSI), Tryptic Soy Agar (TSA) or Nutrient Agar.
* Agglutination tests involve using polyvalent and monovalent antisera.
Salmonella Microbiology
Evolution and Ecological Niche
According to scientific studies, Salmonella has evolved (from E. coli) for over 150 million years through genetic alterations, which has resulted in changes in pathogen ecology.
By evolving into a complex group composed of well over 2300 genetically/phenotypically diverse serotypes, the bacteria has been shown to be capable of infecting a wide range of hosts (both vertebrates and invertebrates).
Over the course of their evolution, Salmonella has also proven to be capable of adapting to and surviving in different habitats in the environment and are therefore referred to as environmental Salmonellae.
The following are some of the ecological niches in which different species and serovars of Salmonella can be found:
Water - Species like S. enteriditis have been shown to survive in water. While various other serotypes have been shown to be able to live and survive in such water bodies as streams, lakes, and river etc, their survival and lifespan is largely dependent on a variety of factors such as temperature, level of oxygen, contamination (animal faces etc.) as well as competing flora among others.
For instance, for some species, reproduction has been shown to be largely favored by warm temperature of the water as well as contamination by animal feces which provided a source of nutrients. In these environments, aquatic fauna such as frogs can act as reservoirs, playing a secondary role in the spread of the organisms.
Sewage - Serovar like S. paratyphi B have been shown to live and multiply in sewage sludge at about 10 degrees Celsius. In one particular study in Sweden, the bacteria could not be attributed to any animal source and was therefore suggested to live freely in this particular niche.
In the event that such sewage is discharged into other environments such as rivers, sea and soil, it can spread and continue to multiply. This is also one method through which they can ultimately infect animals and human beings.
Birds and wild animals - The levels of Salmonella carried by different birds have been shown to vary. For instance, whereas free-living pigeons carry about 17 percent of Salmonella (S. typhimurium) sport and breeding pigeons have been shown to carry much higher levels of the bacteria while wild ducks were found to carry much lower levels.
While transmission to humans from some birds has been reported, most of the birds, including gulls have largely been shown to play a vector role, transferring the bacteria from one site to the next.
Wild and zoo animals have also proven to be sources of exotic and rare serovars. The carrying rate of these animals is largely dependant on the type of animal and their habitats. In reptiles, diseases are not frequently reported as is the case with some birds and wild animals.
* Agricultural and domestic animals have been shown to contribute in contamination especially through the human food chain. Given their exposure to the bacteria in their environment, poultry have been shown to be a significant source of Salmonella.
Salmonella species can also be found in:
- Animal feeds
- Dairy foods
- Aquatic flora
* Salmonella survive in simple celled organisms like amoeba. In these organisms, the bacteria uses a secretion system to protect itself from enzymes that can degrade it.
Metabolism
Salmonella bacteria are facultative anaerobes that are capable of fermenting glucose, mannitol, and sorbotol.
As such, a majority of Salmonella bacteria have the following characteristics:
- Can grow aerobically or anaerobically - This means that they can also grow in the presence of oxygen. While they are capable of using oxygen for respiration, they can also survive through anaerobic respiration by fermenting organic compounds. Here, however, the fermentative pathway is the final electron acceptor in the process.
- Prefer using oxygen for a greater yield of energy during respiration - As such, a majority of Salmonella thrive when oxygen is present. However, studies have shown that in the presence of readily fermentable substances, a good number of these bacteria will refer fermentation. In this case, the sugars have been shown to repress respiratory enzymes, which in turn favors fermentation while minimizing respiration. In the absence of these sugars, as well as other non-fermentable substances, there is an increase of respiratory enzymes that enhances respiration.
- Compared to anaerobiosis, the breakdown rate of sugar during aerobiosis is smaller.
* While a majority of Salmonella bacteria ferment glucose, mannitol and sorbotol, S. arizonae are capable of fermenting lactose.
* The fermentation of sugars by Salmonella results in the production of acids or gas.
Salmonella are also catalase positive and oxidase negative, characteristics that are also used for the purposes of determining the presence of the bacteria.
Some of the other characteristics used to identify the presence of the bacteria in biochemical tests/reactions include:
- Does not hydrolyze urea
- H 2 S positive
- Reduce nitrate to nitrite
- Lysine-Decarboxylase positive
- Arginine-Dihydrolase variable
- Voges-Proskauer positive
* Catalase is an important enzyme found in Salmonella (as well as all other living organisms that are exposed to oxygen). In these organisms, the enzymes are involved in the decomposition of hydrogen peroxide to produce oxygen and water. This helps protect against the cell oxidative damage.
Salmonella Infection
Naturally, infection is acquired through ingestion of water or foods contaminated with the bacteria. However, it may also be acquired through contact with any of the carriers mentioned above.
With over 2500 serovars of Salmonella identified today, more than 1500 have been shown to belong to subspecies enterica. This group is also composed of the majority of bacteria that infect different types of hosts.
Different types of Salmonella affect different hosts, which has led members of the subspecies to be divided into three major groups based on the type of host they infect (wide host specificity).
Specifically:
Unrestricted serovars - This group includes serovars S. typhimurium and S. enteritidis that infects human beings, poultry, swine, mouse, and cattle.
Infections caused by these organisms:
- Enterocolitis (human beings)
- Septicemia in mice
- Asymptomatic in cattle and poultry
Host adapted - includes serovars S. dublin and S. cholerasuis. Hosts for these bacteria include cattle, pigs, chicken, mouse and rarely infect human beings.
Infections include:
- Enterocolitis in cattle (as well as septicemia)
- Fatal systemic infections in swine
- Bacteremia in human beings as well as in mice
Host restricted - This group includes serovars S. typhi, S. gallinarum, and S. abortusequi. These bacteria are found in horses, human beings and poultry.
Infections in the host include:
- Typhus
- Diarrhea
- Septice
* With regards to host specificity, typhi and paratyphi serovars only cause diseases in human beings.
Virulence Factors (Physiology)
Apart from host specificity, several other factors play an important role in the successful infection of the host.
- Endotoxin - Like many other Gram-negative bacteria, some species of Salmonella like Salmonella typhi produce endotoxin (lipopolysaccharides (LPS)), which is a toxic substance that is produced when the outer membrane of the organism is disrupted. This enhances Salmonella infection and inflammation in the affected site.
- Capsule - S. typhi and various strains of S. paratyphi have been shown to possess a capsule as the outermost envelope. These capsules play an important role in the survival of the bacteria given that they cannot be easily removed. As a result, they enhance the infection and have even been associated with resistance to treatment in some cases. Apart from capsules, Salmonella has also been shown to produce outer membrane proteins that allow them to survive in macrophages.
- Adhesives - Apart from capsules that protect the bacteria and enhance their survival, some Salmonella produce such adhesives as fimbrials (and non-fimbrial) that allow the bacteria to remain attached to the surfaces of the infected site. Through reinforced attachment (through these adhesives) bacterial infection is enhanced.
- Flagella - Given that Salmonella tends to affect the intestinal tract, flagella help them move through the intestinal mucus from one site to another.
Human Infections
In human beings and some animals, Salmonella is of clinical importance as they cause salmonellosis and enteric fever. Salmonella infection (salmonellosis) often results in diarrhea, fever (because of inflammation) as well as abdominal cramps that occur 1 to 3 days after infection.
While the infection can last a few days and pass without any treatment, some instances, particularly severe diarrhea, require treatment.
Some infection can spread to the blood stream resulting in toxic shock that can cause death if not treated.
Those at high risk of this infection include:
- Young Children
- Patients with poor/compromised immunity
- Older adults
* Also known as typhoid fever, enteric fever is caused by S. typhi and S. paratyphi and presents the following symptoms:
- General weakness
- Headache
- Cough
- Loss appetite
- Diarrhea and constipation
Pathogenesis
The process through which Salmonella infects and affects human beings in salmonellosis is different from enteric fever.
In salmonellosis, infection starts with ingestion of the bacteria followed by colonization of the lower intestines. This colonization is followed by the invasion of the mucosal layer where acute inflammation takes place.
With regards to enteric fever, oral infection is followed by two weeks of an incubation period after which the bacteria gets to the mucosa layer through the M cells. The infection then proceeds to the local macrophages that transport the bacteria to the mesentarial lymph nodes.
Ultimately, the bacteria can spread to such organs as the lung and liver among others resulting in such complications as empyema, cystitis and typhoid hepatitis.
Prevention and Treatment
For the most part, Salmonella infections are the result of drinking contaminated water, eating uncooked meat, poultry and seafood, uncooked eggs and contaminated fruits and vegetables. It can also be caused through contact with certain pets such as amphibians and reptiles.
Most infections can be prevented by minimizing contact with such pets, particularly for children as well as good hygiene. Here, hygiene involves washing hands with water and soap before eating (and generally keeping them clean), washing all foods before cooking as well as sufficiently cooking meat products that potentially carry the bacteria as well as washing hands after touching pets. These are important prevention tips to avoid infection and the spreading of Salmonella infections.
In some cases, treatment is required. In such cases, treatment involves the use of antibiotics and antimobility drugs as well as replacing fluids and electrolytes. Such drugs as loperamide are used for the purposes of relieving cramping among patients. However, this has been associated with such side effects as diarrhea resulting from the infection.
Microscopy with Gram Stain
A Salmonella sample can be obtained directly from the patient (feces) or from contaminated water/foods. The bacteria may be cultured first using the appropriate agar/media to increase the number of cells.
Sample Preparation
Requirements
- Clean glass slide
- Heat (Bunsen burner)
- Gram stain reagents
- Staining rack
- Sample
Procedure
- Prepare a smear at the center of the glass slide using a cotton swab stick or wire loop. Make sure that the slide, sample and the cotton swab/wire loop are clean to prevent contamination
- Air dry the slide and heat fix (passing over Bunsen flame approximately 3 times and avoid overheating)
- Place the slide on a staining rack and add a few drops of crystal violet onto the sample, gently wash with water
- Add a few drops of Gram iodine (mordant) for between 30 seconds and 1 minute, gently wash with water
- Add a few drops of alcohol (95% alcohol) for about 10 seconds, gently wash with water
- Add a few drops of safranin (counterstain) and rinse with water
- Use a tissue paper to remove excess water by touching the sides of the slide
- View the slide under the microscope starting with lower power
Observation
When viewed under the microscope, Salmonella bacteria, such as Salmonella newport will appear as pink rods. This indicates that it is a Gram-negative bacteria.
Return to Unicellular Organisms
See also Eubacteria page, closely related to E. Coli bacteria and also see Coliform
See also the Prokaryotes main page
Return to Bacteria under the Microscope main page
Return from Salmonella to MicroscopeMaster home
References
C.J. Murray. Salmonellae in the environment. Rev. sci. tech. Off. int. Epiz., 1991, 10 (3), 765-785.
Hiyoshi et al. Typhoidal Salmonella serovars: ecological opportunity and the evolution of a new pathovar. Volume 42, Issue 4, 1 July 2018, Pages 527–541.
Shen, Y. Zhang, Food Microbiology Laboratory for the Food Science Student. Springer International Publishing AG 2017.
Links
https://www.ncbi.nlm.nih.gov/books/NBK8435/
https://www.cdc.gov/salmonella/general/index.html
Find out how to advertise on MicroscopeMaster!
![Salmonella typhimurium Photo: Volker Brinkmann, Max Planck Institute for Infection Biology, Berlin, Germany [CC BY 2.5 (https://creativecommons.org/licenses/by/2.5)], via Wikimedia Commons Salmonella typhimurium Photo: Volker Brinkmann, Max Planck Institute for Infection Biology, Berlin, Germany [CC BY 2.5 (https://creativecommons.org/licenses/by/2.5)], via Wikimedia Commons](https://www.microscopemaster.com/images/256px-Salmonella_typhimurium.png)